Molecular Modeling of Protein-DNA Interactions
Michelle McCully, Biology
Software: NAMD, VMD, ilmm; Hardware: GPU-parallelized and CPU-parallelized nodes
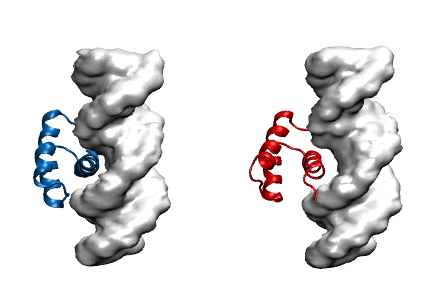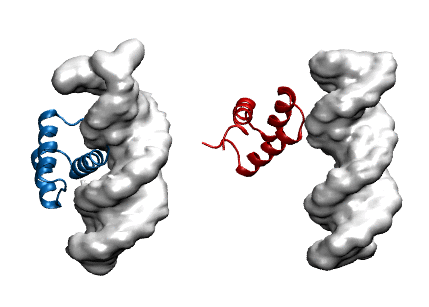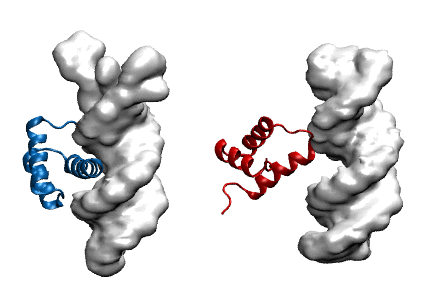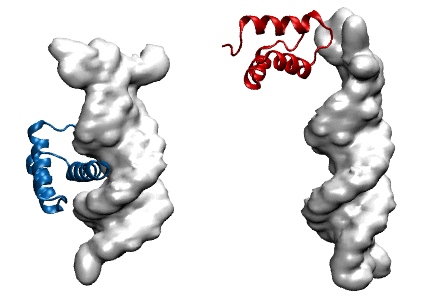
Protein modeling uses computational models to visualize protein structure and dynamics to predict and describe how they function. Proteins are large molecules made in all organisms and perform countless jobs in cells. The McCully Lab is particularly interested in the differences between the proteins that organisms make naturally and proteins that scientists design in the lab. They use a combination of experimental, computational, and theoretical techniques to investigate the relationships between these proteins’ shapes, movements, and functions. This work allows scientists to understand the mechanisms by which proteins maintain their functions in harsh environments, for example industrial settings and long-term storage. Undergraduate student researchers use the WAVE facilities to perform molecular dynamics simulations of protein-DNA complexes in order to predict how well they bind. In the image, the blue protein stays bound to DNA (gray) in their simulations, whereas the red one does not.